A few months ago I gave a presentation in the Connections: Life Sciences & Healthcare virtual event. It was about building a Knowledge Graph using public RDF resources. You can watch the recording here or even reproduce the whole session following the instructions in this repository.
I went through the content again recently and I found one particular bit of that session that was specially interesting and worth spending a QuickGraph on. I’m talking of course of the reconciliation of taxonomies. Let’s dive in.
Building the graph
We are loading three taxonomies of diseases, one from Wikidata, one from MeSH and finally the one from the DiseaseOntology. Both the Wikidata and the DiseaseOntology include cross-references to the other two. We will load these cross references too as we will use them to reconcile the three taxonomies.
The target model is very straightforward. Taxonomy categories (diseases) are labelled based on their provenance: WD_Disease, Mesh_Disease and DO_Disease and linked to their parent categories through PARENT_OF relationships. The cross references between categories are represented with SAME_AS relationships.
Probably worth mentioning that the model we are building for this QG is slightly different from the one I created for the Connections: Life Sciences & Healthcare as we are putting the focus on the taxonomy reconciliation here.
Before we start…
Since we are going to be using public RDF resources, the process will inevitably start by making sure we have the n10s plugin installed and the graph properly configured. Following the instructions in the manual we will first create a uniqueness constraint on Resource(uri):
CREATE CONSTRAINT n10s_unique_uri ON (r:Resource)
ASSERT r.uri IS UNIQUE;And then a graph configuration with the following parameters:
call n10s.graphconfig.init({ handleVocabUris: "IGNORE", classLabel: "DO_Disease", subClassOfRel: "HAS_PARENT"})Refer to the manual for any additional settings.
Loading a taxonomy of infectious diseases from Wikidata.
The following SPARQL query gets what we need. The key element is the pattern ?dis wdt:P31/wdt:P279* wd:Q18123741 that explores recursively the Instance of (P31) and subclass of (P279) relationships down from the concept Infectious Disease (Q18123741). The query also returns -when present- the references to the equivalent MeSH (P486) and DiseaseOntology(P699) concepts.
PREFIX neo: <neo://voc#>
construct {
?dis a neo:WD_Disease ;
neo:label ?disName ;
neo:HAS_PARENT ?parentDisease ;
neo:SAME_AS ?meshUri ;
neo:SAME_AS ?diseaseOntoUri .
}
where {
?dis wdt:P31/wdt:P279* wd:Q18123741 ;
rdfs:label ?disName . filter(lang(?disName) = "en")
optional { ?dis wdt:P279 ?parentDisease .
?parentDisease wdt:P31/wdt:P279* wd:Q18123741 }
optional { ?dis wdt:P486 ?meshCode . bind(URI(concat("http://id.nlm.nih.gov/mesh/",?meshCode)) as ?meshUri) }
optional { ?dis wdt:P699 ?diseaseOntoId . bind(URI(concat("http://purl.obolibrary.org/obo/",REPLACE(?diseaseOntoId, ":", "_"))) as ?diseaseOntoUri) }
}It is quite straightforward to embed the SPARQL query in the rdf.import procedure. This procedure sends an HTTP request to the Wikidata SPARQL endpoint (http://query.wikidata.org/sparql) and gets the returned triples imported into Neo4j:
WITH '...previous Wikidata SPARQL fragment...' AS query
CALL n10s.rdf.import.fetch(
"https://query.wikidata.org/sparql?query=" + apoc.text.urlencode(query),
"N-Triples",
{ headerParams: { Accept: "text/plain"}})
YIELD terminationStatus, triplesLoaded, triplesParsed, namespaces, extraInfo
RETURN terminationStatus, triplesLoaded, triplesParsed, namespaces, extraInfoIf everything goes well, two and a half thousand triples should be imported:
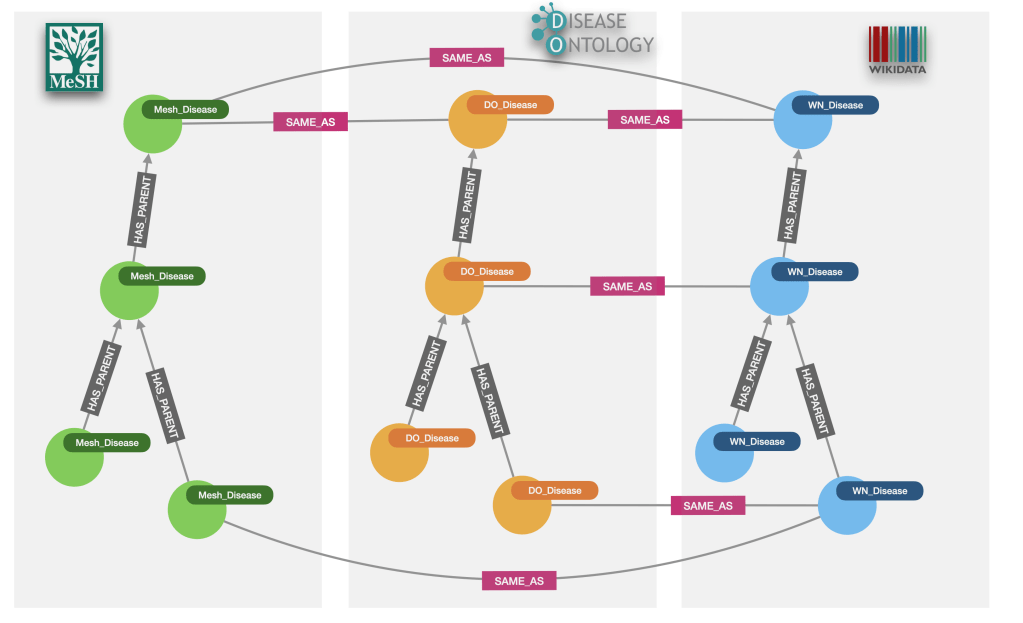
And our graph should now contain the first taxonomy of diseases linked through HAS_PARENT relationships and with optional references to both MeSH descriptors and the ids from the DiseaseOntology. Here is an example:
Note that the blue nodes only have a uri and will be completed when we import the other taxonomies.
Wikidata is manually curated so it’s not unusual to find ‘shortcuts’ in taxonomies. In our case these are redundant HAS_PARENT relationships, where a category is connected to its immediate super-category (expected), but also to some other ancestor like in the diagram below.
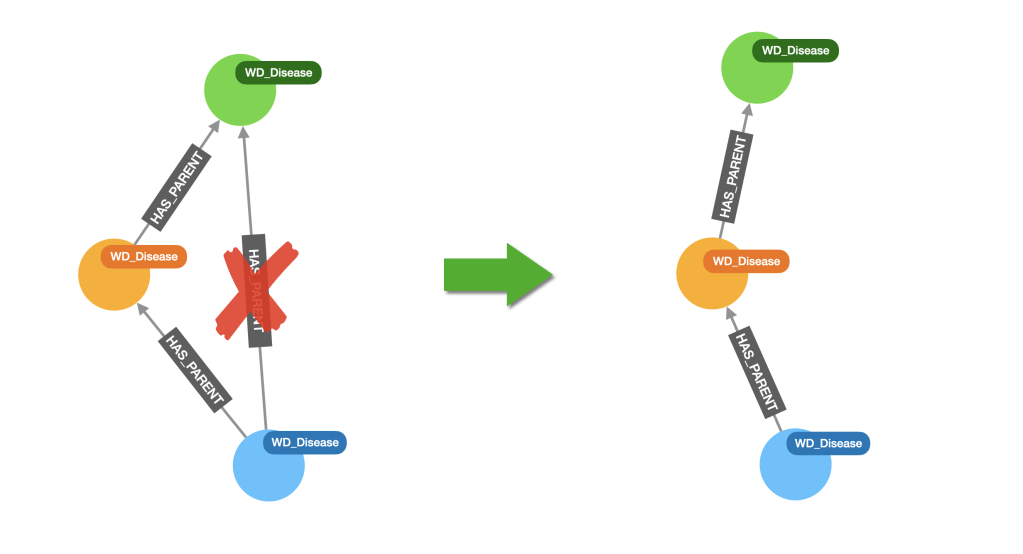
These links are not needed and can complicate the analysis we want to carry out. The good news is they can be easily removed with this simple cypher query:
MATCH (v:WD_Disease)<-[co:HAS_PARENT*2..]-(child)-[shortcut:HAS_PARENT]->(v) DELETE shortcutLoading a taxonomy of diseases from MeSH
Just like in the previous case, we built a SPARQL query on the MeSH controlled vocabulary that returns the subtaxonomy of infectious diseases. It navigates the meshv:broaderDescriptor relationship down from the mesh:D007239 descriptor representing Infections.
PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
PREFIX meshv: <http://id.nlm.nih.gov/mesh/vocab#>
PREFIX mesh: <http://id.nlm.nih.gov/mesh/>
PREFIX neo: <neo://voc#>
CONSTRUCT {
?s a neo:Mesh_Disease;
neo:label ?name ;
neo:HAS_PARENT ?parentDescriptor .
}
FROM <http://id.nlm.nih.gov/mesh>
WHERE {
{
?s meshv:broaderDescriptor* mesh:D007239
}
?s rdfs:label ?name .
optional {
?s meshv:broaderDescriptor ?parentDescriptor .
}
}We embed the SPARQL into a rdf.import to issue the query to the MeSH SPARQL endpoint (http://id.nlm.nih.gov/mesh/sparql) and get the returned triples imported.
WITH '...previous MeSH SPARQL fragment...' AS query
CALL n10s.rdf.import.fetch(
"https://id.nlm.nih.gov/mesh/sparql?format=TURTLE&query=" + apoc.text.urlencode(query),
"Turtle")
YIELD terminationStatus, triplesLoaded, triplesParsed, namespaces, extraInfo
RETURN terminationStatus, triplesLoaded, triplesParsed, namespaces, extraInfo
All going well, just under three thousand triples should be imported:
And the imported taxonomy should link automagically with the Wikidata one. Neosemantics takes care of merging the nodes with the same uri so we should now be able to find in our graph parallel fragments of the Wikidata and MeSH taxonomies.
We will also do the usual taxonomy cleansing on the MeSH side and remove the redundant HAS_PARENT links. Same cypher fragment, now applied to Mesh_Disease nodes:
MATCH (v:Mesh_Disease)<-[co:HAS_PARENT*2..]-(child)-[shortcut:HAS_PARENT]->(v)
DELETE shortcutThis is what the fragment queried before looks like now:
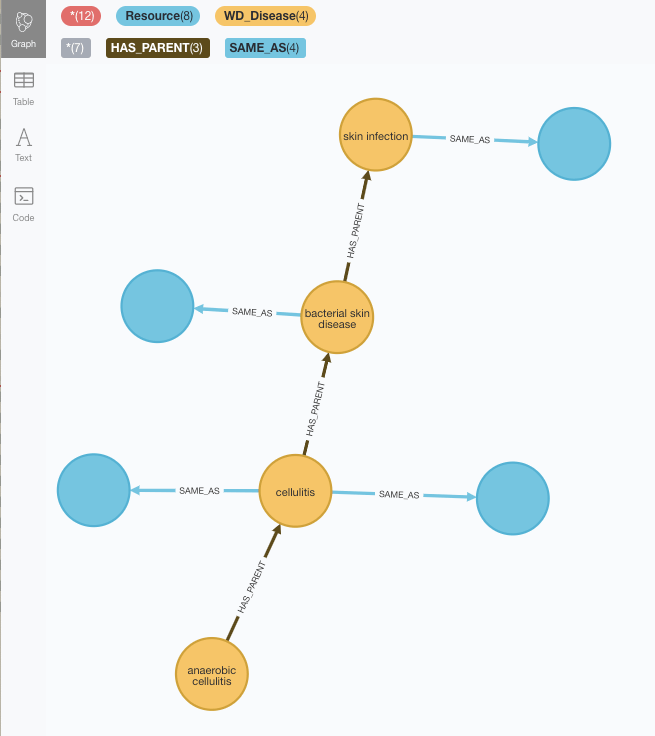
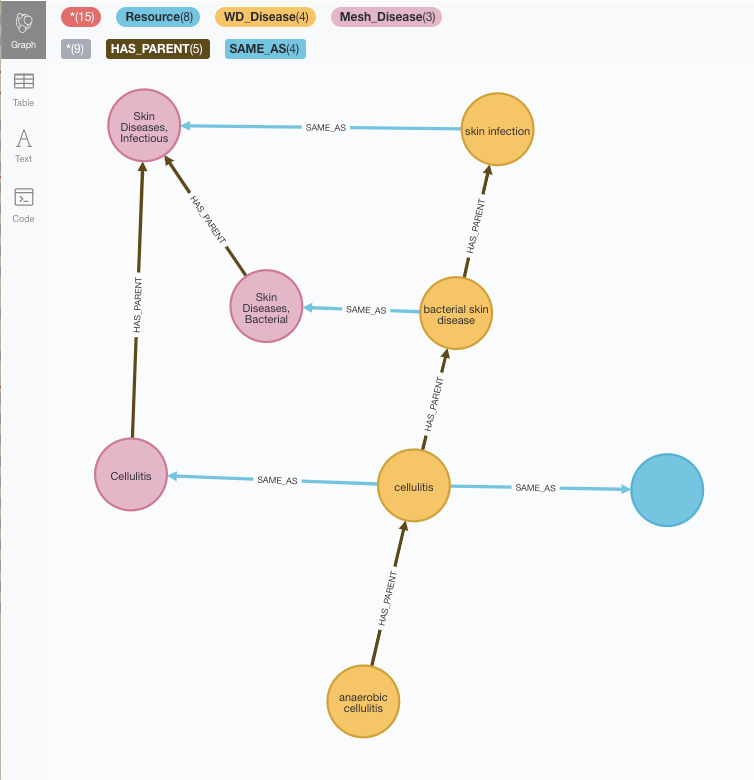
The first thing we notice is that yes, some of the placeholders have been filled with the MeSH taxonomy elements although some are still unlinked (the blue one to the right is a reference to an element in the Disease Ontology which has not been loaded yet).
The other thing we notice is that the taxonomies are not identical, and sequences that are lineal in Wikidata are not in MeSH. Well, that’s exactly what we were after in our Taxonomy reconciliation exercise, so here we are.
Loading the Human Disease Ontology
Wit the Human Disease Ontology we are going to follow a slightly different approach simply because we can access it in a single file instead of having to extract it via SPARQL from a triple store.
We will invoke the onto.import procedure passing the url of the file in the OBO Foundry.
call n10s.onto.import.fetch("http://purl.obolibrary.org/obo/doid.owl","RDF/XML");The onto.import method only imports the classes and relationships and their hierarchical relationships so we need to apply a second pass to get the cross references too. The rdf.stream method will do the job for us. We just need to select the triples having hasDbXref as predicate and then import those where the object starts with “MESH:”. The Disease Ontology represents the cross references with literal values containing the ids of objects in other taxonomies.
call n10s.rdf.stream.fetch("http://purl.obolibrary.org/obo/doid.owl","RDF/XML", { limit : 999999}) yield subject, predicate, object
where predicate = "http://www.geneontology.org/formats/oboInOwl#hasDbXref" and object starts with "MESH:"
MATCH (doe:Resource { uri: subject}),
(mesh:Resource { uri: "http://id.nlm.nih.gov/mesh/" + substring(object,5)})
MERGE (doe)-[:SAME_AS]->(mesh);And we have a complete graph now! Our three taxonomies will sometimes align nicely, both structurally and also from the point of view of the cross-referencing, like in the next image for the subcategories of ‘Coccidiosis’, but this will not always be the case as we already anticipated in the previous section.
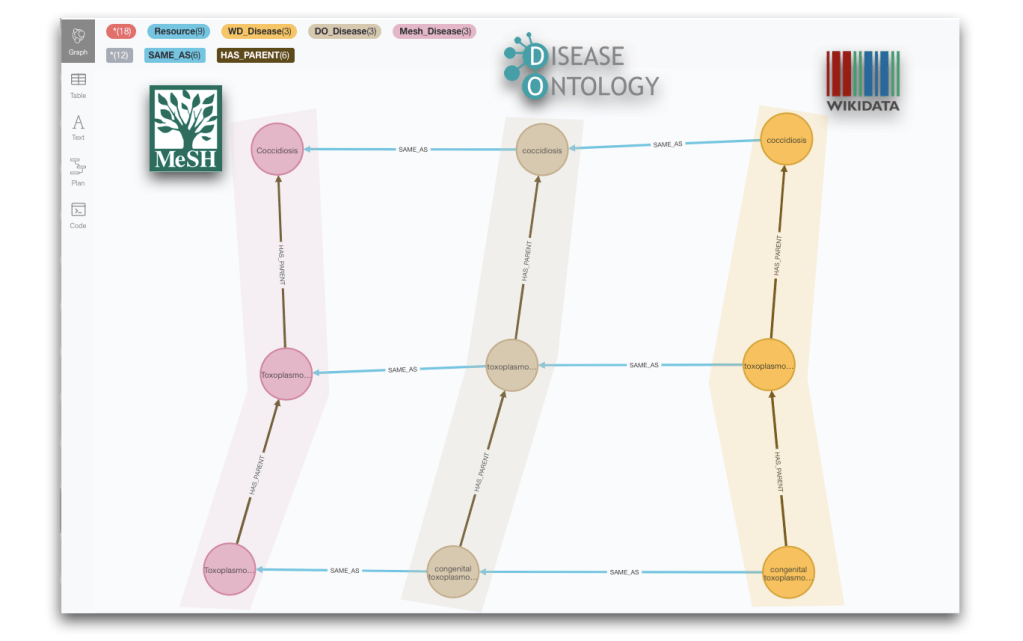
Taxonomy analysis and reconciliation
In order to keep this post a QuickGraph 🙂 we will present at some ideas on how to compare the taxonomies pairwise and all together but I’ll leave it up to you to try other patterns (there are many more!) and contribute them to the repository (link coming soon).
Detecting different granularities in taxonomies (pairwise comparison)
We can detect situations where a taxonomy has a more granular classification than other by finding two points where the taxonomies meet (top and bottom) and finding for each of them HAS_PARENT paths of different lenghts. This query would return such cases between the Disease Ontology and MeSH:
MATCH topLink = (topDo:DO_Disease)-[:SAME_AS]-(topMesh:Mesh_Disease)
MATCH bottomLink = (bottomDo:DO_Disease)-[:SAME_AS]-(bottomMesh:Mesh_Disease)
MATCH txnDo = (topDo)<-[:HAS_PARENT*]-(bottomDo)
MATCH txnMesh = (topMesh)<-[:HAS_PARENT*]-(bottomMesh)
WHERE length(txnDo) <> length(txnMesh)
RETURN * limit 10It will return examples like this one:
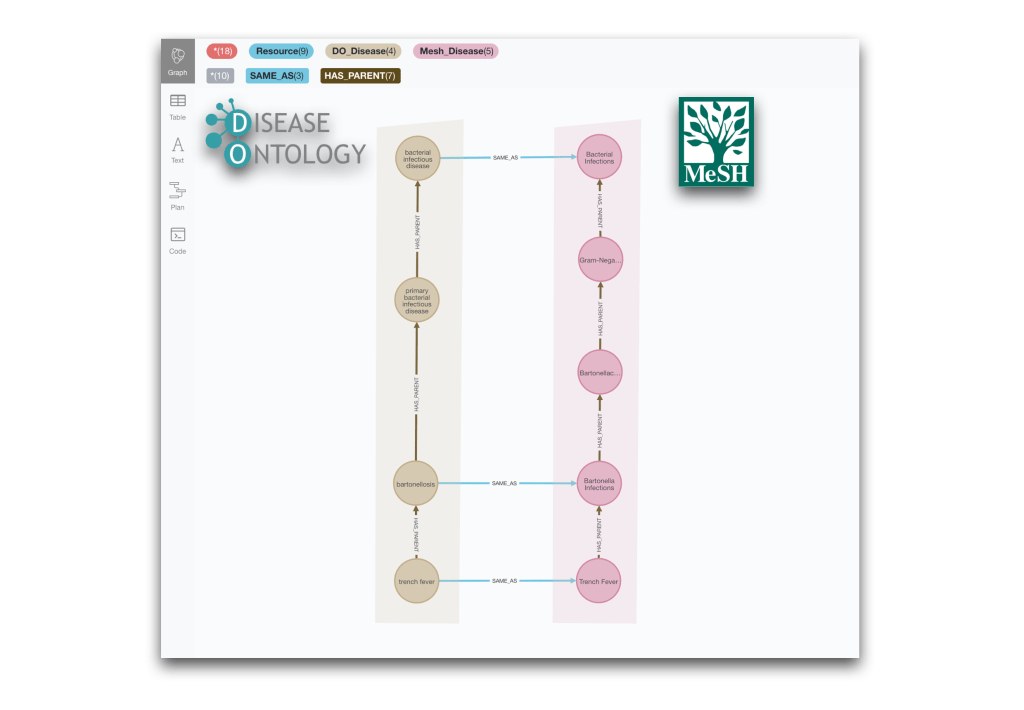
We can detect the same situation between the Wikidata and the Disease Ontology taxonomies by just modifying the types of the nodes.
profile MATCH topLink = (topW:WD_Disease)-[:SAME_AS]-(topDo:DO_Disease)
MATCH bottomLink = (bottomW:WD_Disease)-[:SAME_AS]-(bottomDo:DO_Disease)
MATCH txnDo = (topDo)<-[:HAS_PARENT*]-(bottomDo)
MATCH txnW = (topW)<-[:HAS_PARENT*]-(bottomW)
WHERE length(txnDo) <> length(txnW)
RETURN * limit 10In this case we are showing multiple cases (taxonomy fragments) in the same visuailsation. Feel free to play with the limit in the previous query to get more results:
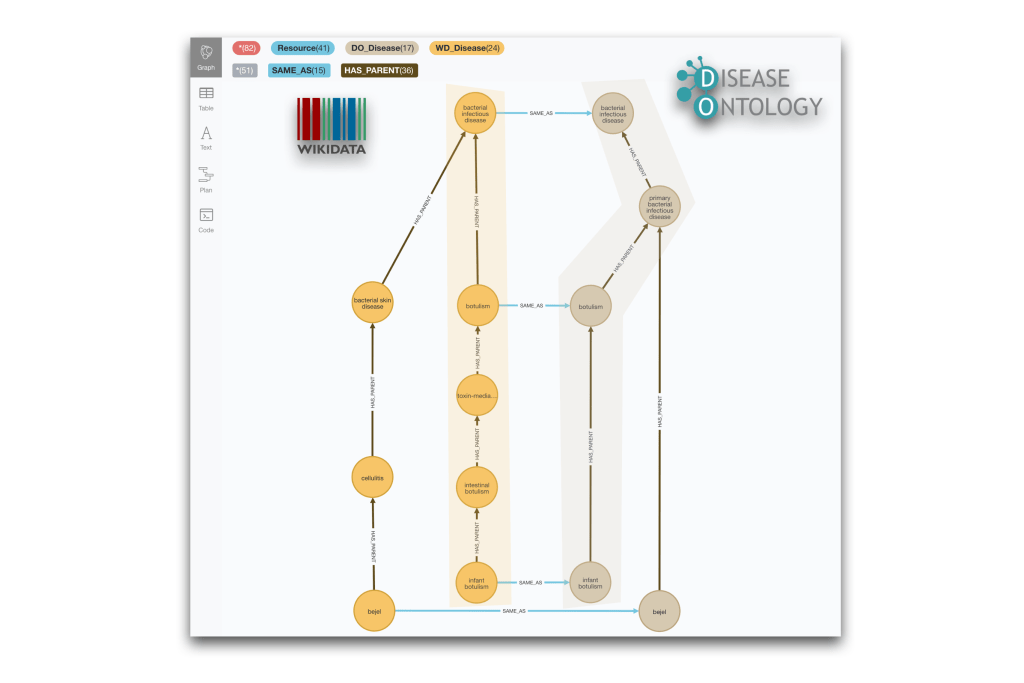
And for completeness we show also an example of different granularity between Wikidata and MeSH. We leave the writing of the query to the interested reader.
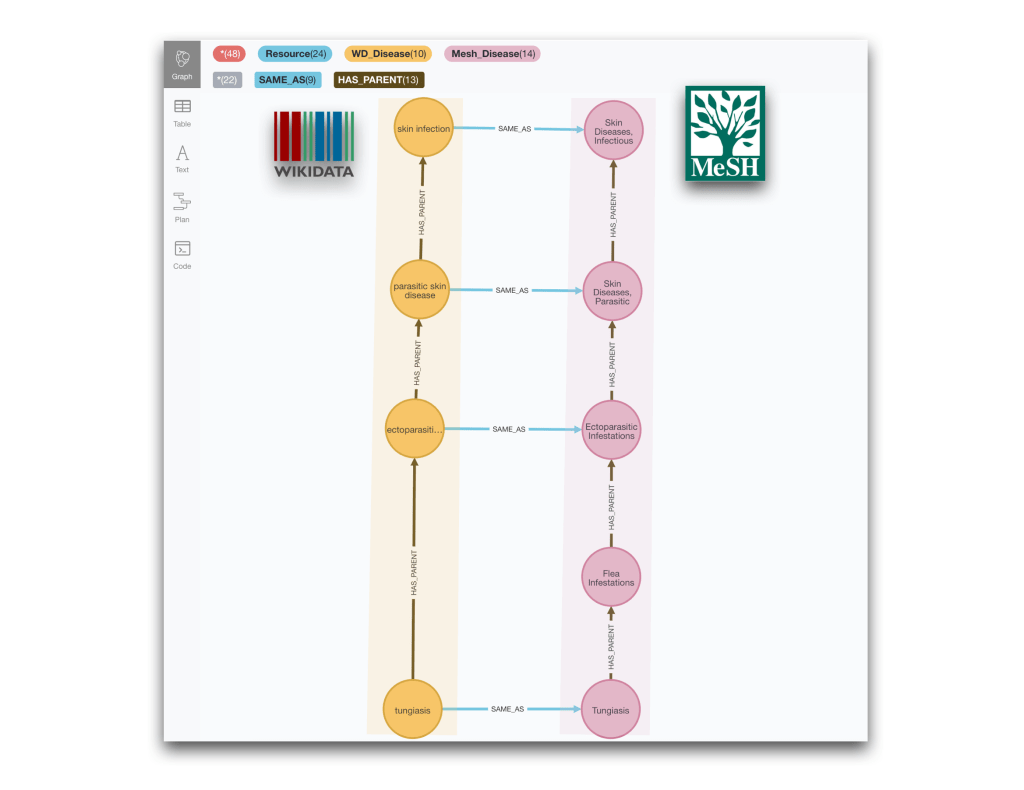
Understanding multiple cross-references
Some categories may include multiple cross-references. As in a category in taxonomy A with multiple equivalent categories in taxonomy B. Since these taxonomies are manually curated, we will of course assume that this was intentional and not an accident.
We can detect which combinations of sources verify this with the following query:
MATCH (x:DO_Disease) WHERE size ((x)-[:SAME_AS]-(:Mesh_Disease)) > 1
RETURN count(x) as ct, "DO-MeSH" as what
UNION
MATCH (x:DO_Disease) WHERE size ((x)-[:SAME_AS]-(:WD_Disease)) > 1
RETURN count(x) as ct, "DO-Wikidata" as what
UNION
MATCH (x:Mesh_Disease) WHERE size ((x)-[:SAME_AS]-(:DO_Disease)) > 1
RETURN count(x) as ct, "MeSH-DO" as what
UNION
MATCH (x:Mesh_Disease) WHERE size ((x)-[:SAME_AS]-(:WD_Disease)) > 1
RETURN count(x) as ct, "MeSH-Wikidata" as what
UNION
MATCH (x:WD_Disease) WHERE size ((x)-[:SAME_AS]-(:Mesh_Disease)) > 1
RETURN count(x) as ct, "Wikidata-MeSH" as what
UNION
MATCH (x:WD_Disease) WHERE size ((x)-[:SAME_AS]-(:DO_Disease)) > 1
RETURN count(x) as ct, "Wikidata-DO" as whatproducing as result:
And now we can analyse each of the cases in detail. For example, for the case of the Disease Ontology, the following query returns all the nodes of type MeSH taxonomy that they match to and also checks whether there is a hierarchical relationship between them.
MATCH multiXRef = (md1:Mesh_Disease)-[:SAME_AS]-(start:DO_Disease)-[:SAME_AS]-(md2:Mesh_Disease)
OPTIONAL MATCH link = (md1)-[r:HAS_PARENT*]->(md2)
RETURN multiXRef, link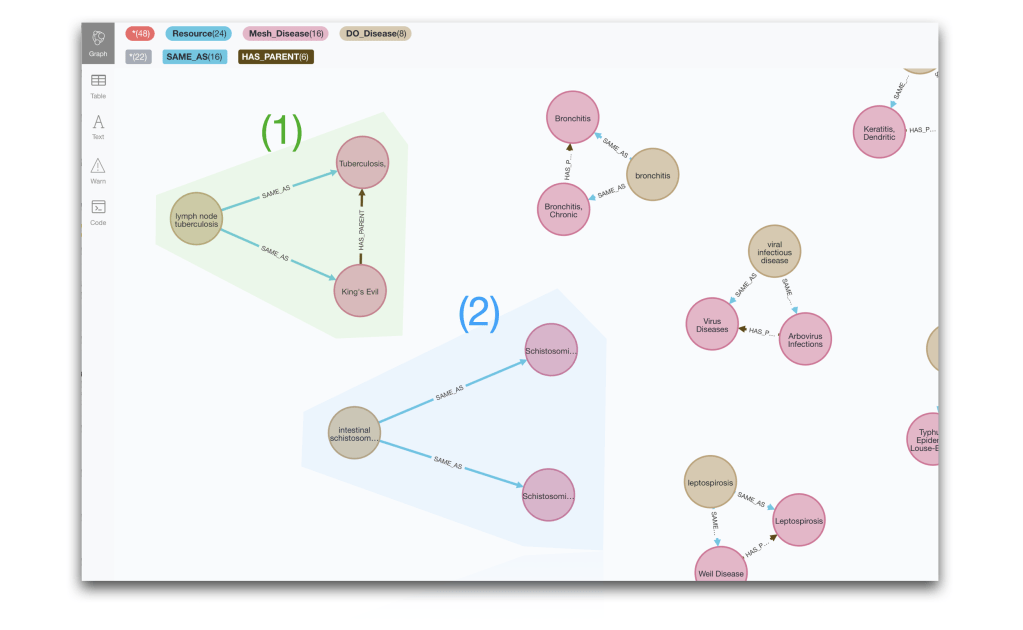
In this post we are focusing on detecting patterns and visualising them. But we could try to implement some actions to harmonise the ontologies, for example for case (1) we could determine that that SAME_AS link to the child category could be removed as it would be subsumed by the SAME_AS to the parent category. In case (2) some human intervention would be required to disambiguate the linkage.
If we run the same type of analysis for the MeSH-DO case, we get a couple more variants (or pseudo variants). Case (1) is the one seen before and (2) is just an aggregation of multiple of type (1). Then case (3) extends the hierarchical relation between the two matched entities to multiple hops. The query producing these results is the following:
MATCH multiXRef = (md1:DO_Disease)-[:SAME_AS]-(start:Mesh_Disease)-[:SAME_AS]-(md2:DO_Disease)
OPTIONAL MATCH link = (md1)-[r:HAS_PARENT*]->(md2)
RETURN multiXRef, link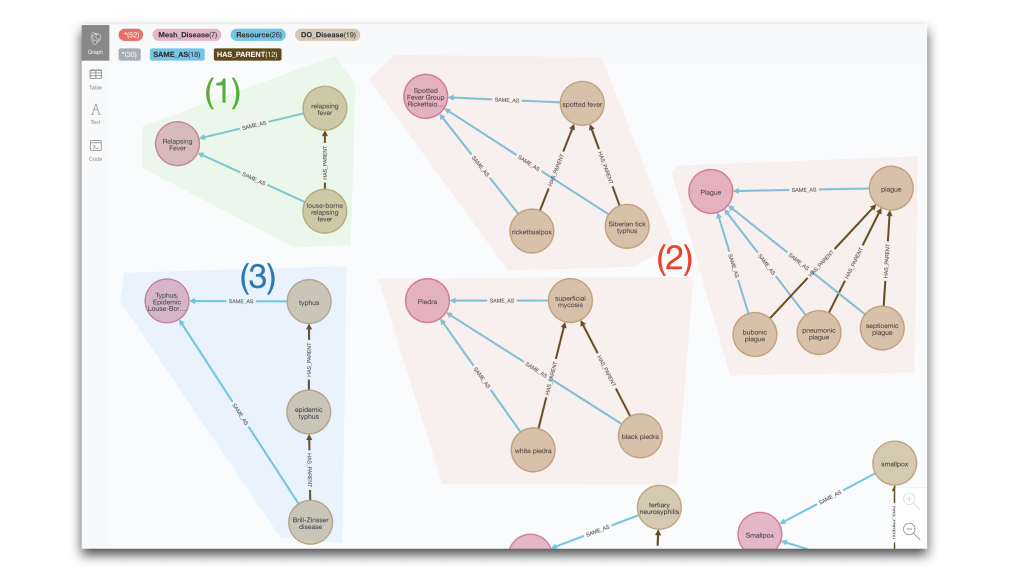
We leave to the interested reader the application of the same analysis to the remaining case (Wikidata-DO).
Triadic closures (aligning the three taxonomies)
In the case of a perfect alignment between the three taxonomies, we should find triangles connecting the matching entities. We can identify these with a simple Cypher pattern:
MATCH triangle = (wdid:WD_Disease)-[:SAME_AS]-(do:DO_Disease)-[:SAME_AS]-(md:Mesh_Disease)-[:SAME_AS]-(wdid)
WHERE size((wdid)-[:SAME_AS]-()) = size((do)-[:SAME_AS]-()) = size((md)-[:SAME_AS]-()) = 2
RETURN count(triangle) limit 50Note that for the matching to be perfect there should be no extra SAME_AS matches in addition to the ones forming the triangle, hence the check on the degree of the nodes.
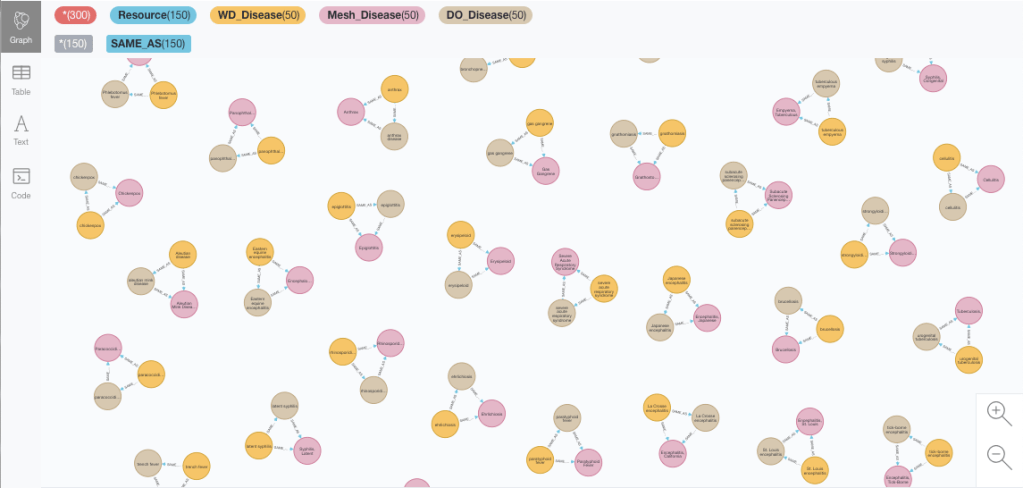
There are 222 triangles (you can get the count by swapping the RETURN clause of the previous query with something like RETURN count(triangle), which represent a third of the entities in the Wikidata taxonomy if we take it as the reference. Not bad. Now let’s look at the not-so-perfect cases.
Triangles missing one leg (side)
The incomplete triangles from the Wikidata point of view could be found with this pattern (note that we may want to relax the condition on the node degree. See previous section):
MATCH incomplete = (wdid:WD_Disease)-[:SAME_AS]-(do:DO_Disease)-[:SAME_AS]-(md:Mesh_Disease)
WHERE NOT (md)-[:SAME_AS]-(wdid) AND size((wdid)-[:SAME_AS]-()) = size((md)-[:SAME_AS]-()) = 1 AND size((do)-[:SAME_AS]-()) = 2
RETURN incomplete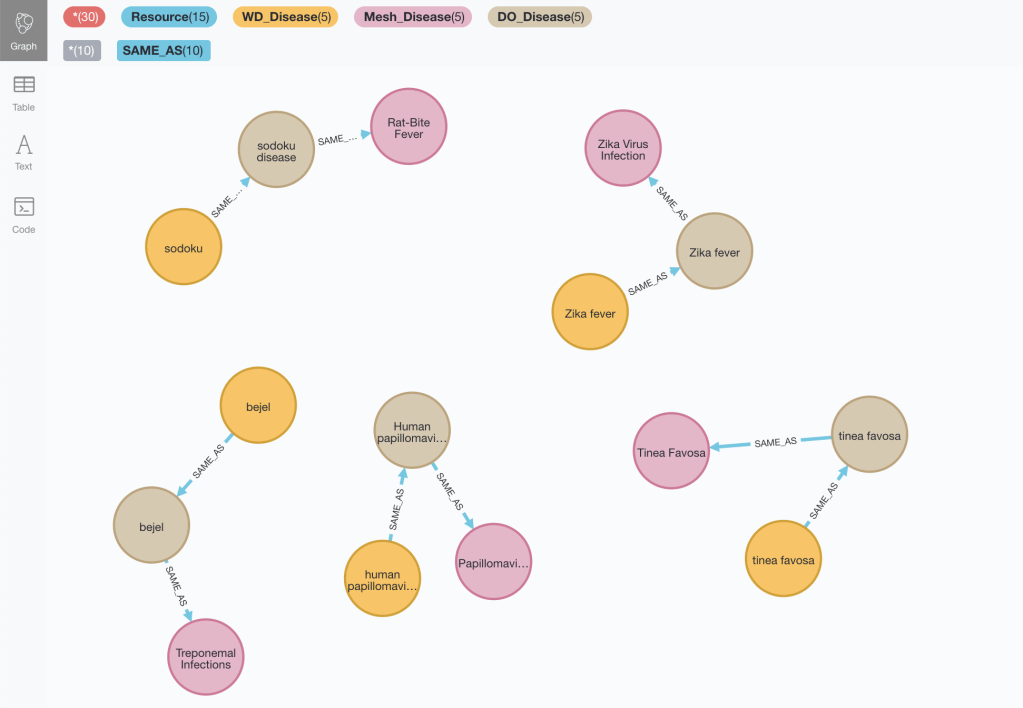
Generating cross-references for Wikidata (link prediction with triangles)
We can use the previous query to ‘predict’ references missing from Wikidata.
MATCH incomplete = (wdid:WD_Disease)-[:SAME_AS]-(do:DO_Disease)-[:SAME_AS]-(md:Mesh_Disease)
WHERE NOT (md)-[:SAME_AS]-(wdid) AND size((wdid)-[:SAME_AS]-()) = size((md)-[:SAME_AS]-()) = 1 AND size((do)-[:SAME_AS]-()) = 2
RETURN wdid.uri as subject, "http://www.wikidata.org/prop/direct/P486" as predicate, n10s.rdf.getIRILocalName(md.uri) as object;Producing the following triples (again, removing the filter on the node degrees more could be generated)
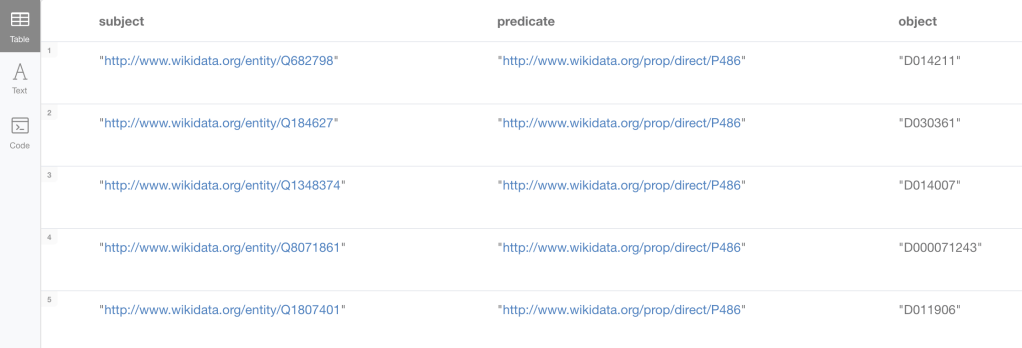
What’s interesting about this QuickGraph?
It highlights the many possible types of misalignments between taxonomies but at the same time, it shows how the ‘shapes’ or patterns we want to detect can be easily expressed using cypher.
Also, we have focused on detecting the patterns and visualising them but we could try to implement some actions to harmonise the ontologies. Maybe the topic of a future post? Or even better, why don’t you share your ideas on this.
As always, give it all a try using the instructions and queries in this post and share your experience or your questions on the neo4j community site.
Thanks for your interest, see you in the next QG.

Thank you for this Quickgraph series !
LikeLiked by 1 person